Current Research Projects
Microbiome & Computational Biology Laboratory
Rumen Microbiome
Project 1: Rumen Hydrogen Sink Mechanisms
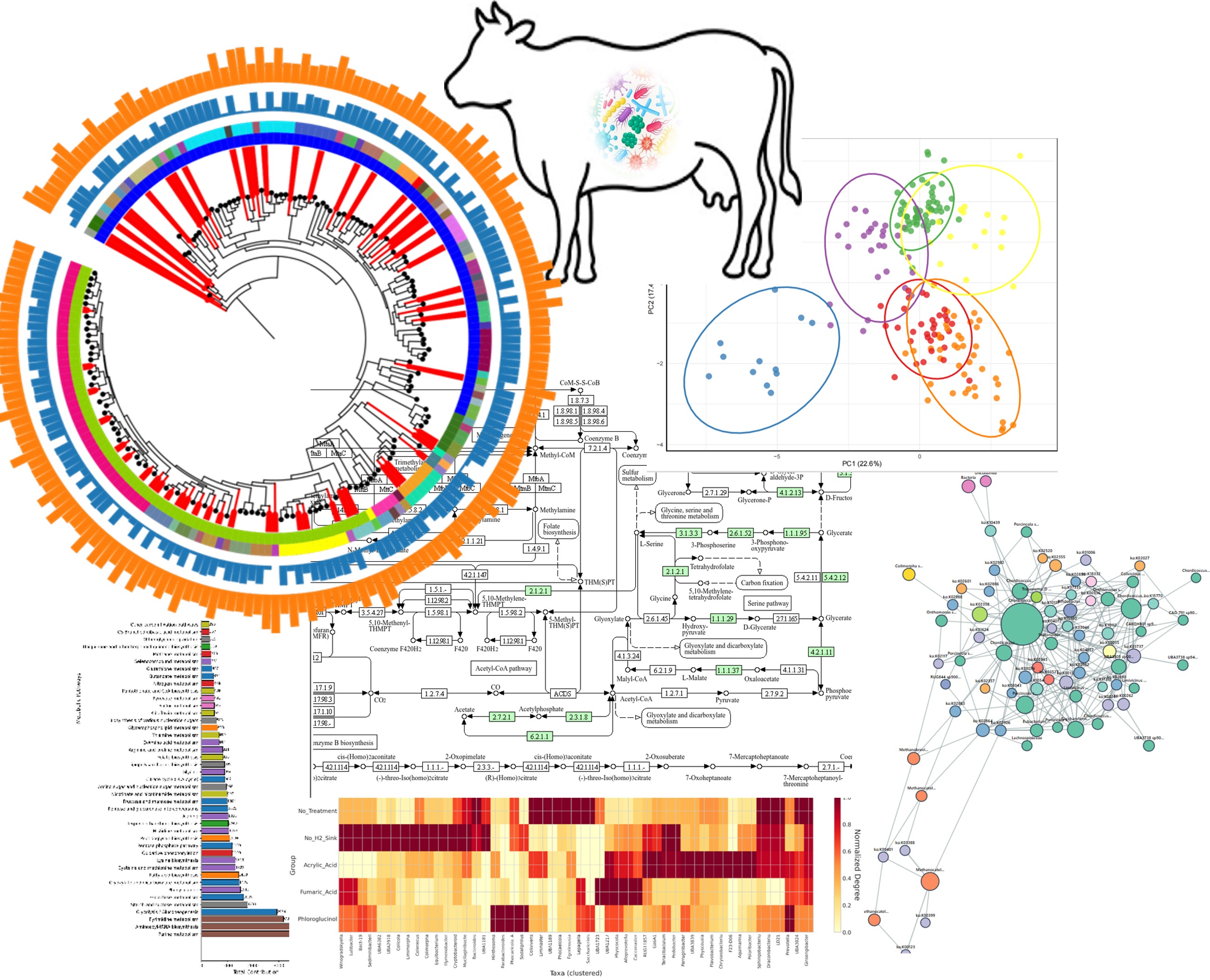
This project investigates alternative hydrogen pathways in the rumen microbiome when methane production is inhibited. Using a carefully designed experimental approach with Holstein dairy cattle, we're studying how the rumen microbial community adapts to different conditions.
- Multi-omics analysis of microbiome adaptation
- Testing of novel compounds for environmental impact
- Controlled experimental design with Holstein cows
- Comprehensive metagenomic and metatranscriptomic analysis
MetaMAG Explorer
Project 2: MetaMAG Explorer Pipeline
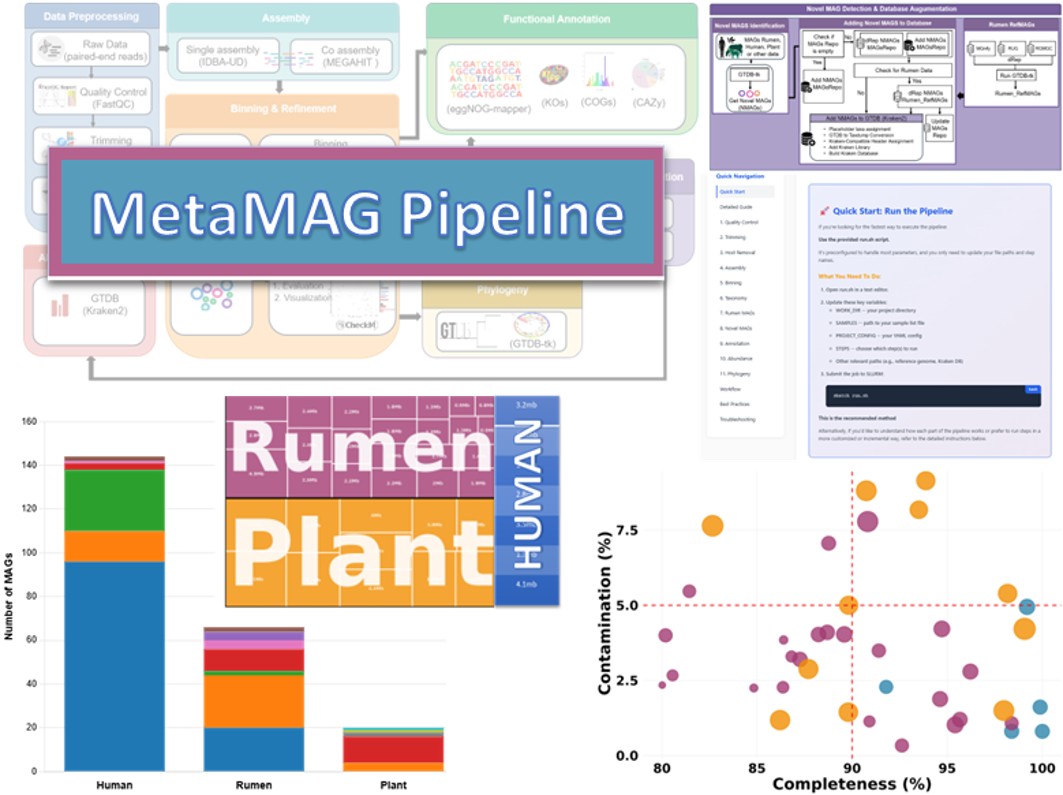

We're developing a computational pipeline for the discovery and validation of novel microbial genomes from metagenomic data. This tool enhances reference databases to improve microbiome analysis across different environments.
- Novel microbial genome discovery workflow
- Database enhancement for improved microbiome analysis
- Validation across diverse ecosystems (Human, Rumen, Plant)
- Integration of multiple bioinformatics tools into a unified pipeline